Lots of the medication we make the most of in fashionable drugs are naturally produced by microbes. Penicillin, an antibiotic derived from sure molds, is likely one of the most notable pure merchandise as a result of its recognition as one of many largest advances in drugs and human well being. As DNA sequencing has turn out to be cheaper and quicker, scientists now have entry to tons of of hundreds of microbial genomes and the pure merchandise they produce.
Nevertheless, Doug Mitchell (MMG), the John and Margaret Witt Professor of Chemistry at College of Illinois, says this pales compared to the variety of compounds these organisms have the capability to make utilizing the genetic pathways they possess.
“That is simply the tip of the iceberg,” mentioned Mitchell. “There is a disparity in what we all know as we speak by way of recognized molecules versus what nature has the capability to provide. Like 100 to 1 no less than.”
One group of pure merchandise that has turn out to be a preferred supply of antibiotics is known as ribosomally synthesized and post-translationally modified peptides, or just, “RiPPs.” Conventional strategies for accessing RiPPs are gradual, and contain taking genes one after the other and placing them right into a mannequin organism, like E. coli, to see what compound it produces.
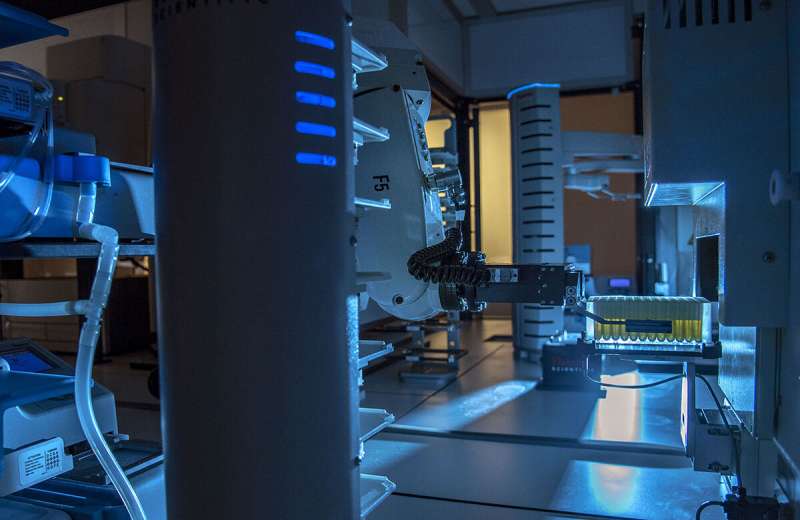
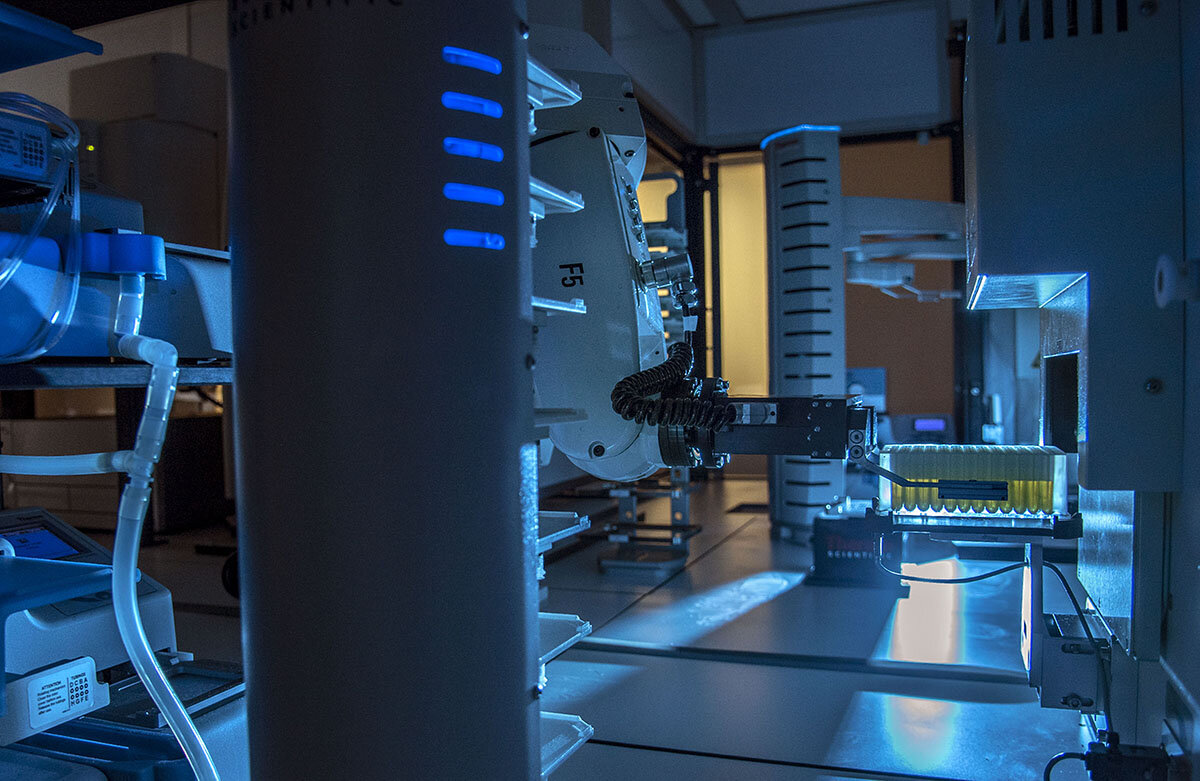
Nevertheless, in a brand new paper ensuing from an enormous collaborative effort on the Carl R. Woese Institute for Genomic Biology, researchers had been in a position to uncover and characterize new RiPPs at an unprecedented pace and scale utilizing the Illinois Organic Foundry for Superior Biomanufacturing (iBioFAB). This can be a laboratory automation system that may consider and assemble a number of artificial gene pathways from tons of of genes without delay, one thing that may historically take many researchers and rather more time to perform.
The mission incorporates a collaboration between Mitchell’s lab, the lab of Huimin Zhao (BSD/GSE chief/CABBI/CGD/MMG), the Steven L. Miller Chair of chemical and biomolecular engineering, and the lab of Wilfred van der Donk (MMG), Richard E. Heckert Endowed Chair in Chemistry and Howard Hughes Medical Institute Investigator.
The three co-first authors, Alex Battiste, fourth-year Ph.D. pupil within the Mitchell lab, Chengyou Shi, fifth-year Ph.D. candidate within the Zhao lab, and Richard Ayikpoe, a postdoc within the van der Donk lab, described how they every led part of the mission of their respective labs. Shi’s crew ordered artificial genes after which assembled them into candidate pathways, or gene clusters, utilizing iBioFAB built-in with a genome mining program known as RODEO. Then, totally different courses of the gene clusters got to Battiste and Ayikpoe’s groups to check which pathways had been useful and prone to produce new RiPPs in E. coli. Any buildings of RiPPs that confirmed antibiotic actions had been characterised intimately by Ayikpoe’s crew. The high-throughput expertise allowed for 96 pathways composed of about 400 genes to be examined without delay, with the manufacturing of 30 new compounds.
“In contrast with conventional RiPP discovery strategies, our platform is scalable and high-throughput in lots of features, from the biosynthetic gene cluster identification, the cloning, the manufacturing, and detection and characterization,” mentioned Shi. “This, I might say, is the primary such platform for giant scale RiPP discovery.”
Out of the brand new compounds found, three had been discovered to have antibacterial properties. When examined towards Klebsiella pneumoniae, that are extremely virulent antibiotic-resistant micro organism, the newly found antibacterial RiPPs had been discovered to be efficient at killing the damaging micro organism. The researchers say this may very well be a brand new avenue for locating compounds which might be efficient towards micro organism which might be immune to present antibiotic medication.
“We discovered three RiPPs which have antimicrobial properties towards pathogens which might be recognized to be concerned in hospital acquired infections, together with Klebsiella,” mentioned Ayikpoe. “This analysis reveals that through the use of this platform to increase the variety of biosynthetic gene clusters that we are able to display screen without delay, we usually tend to uncover anti-microbial compounds that might have therapeutic properties.”
The crew says the aim of the paper is two-fold: to exhibit the flexibility of the high-throughput expertise to rapidly assemble and check gene clusters for brand new RiPPs, and in addition to emphasise the type of large-scale collaborative initiatives which might be made potential throughout the IGB. “There is no means that anyone of our labs might have performed all of this on their very own. The IGB has supplied the crucible for this sort of interdisciplinary analysis,” Mitchell mentioned.
Battiste described how the IGB evokes collaborative initiatives like this one naturally by way of its design. “The IGB makes it very simple to only discuss to individuals while you see them on a regular basis in your theme, which lowers the barrier for beginning initiatives with them,” Battiste mentioned. “Everybody within the MMG theme works on related stuff even when we’re from totally different labs. So all of us have several types of experience however they mesh properly collectively, and also you get to study in regards to the kinds of methods they’re utilizing. It has been considered one of my favourite elements of working right here, the sense of camaraderie amongst all the individuals on the crew.”
To focus on the spirit of collaboration embodied by their paper, the labs are working with the Division of Chemistry to create a video to showcase each their analysis and all that the IGB provides to empower initiatives like these, and to hopefully encourage extra of them. The video is ready to launch quickly to accompany the publication of the paper in Nature Communications.
All three co-first authors described how their training, analysis, and job prospects have benefitted enormously from their time on the IGB, highlighting that it’s each the individuals and the expertise collectively that make IGB a terrific place to conduct analysis. “The collaborative ambiance that the IGB provides in variety and development, each by way of science and social life, is actually exceptional.” mentioned Ayikpoe.
New instrument, RODEO, guarantees to seize the breadth of microbial biosynthetic potential
Richard S. Ayikpoe et al, A scalable platform to find antimicrobials of ribosomal origin, Nature Communications (2022). DOI: 10.1038/s41467-022-33890-w
College of Illinois at Urbana-Champaign
Quotation:
Collaborative crew discovers new pure merchandise, for use as sources of antibiotics, at unprecedented pace (2022, October 18)
retrieved 18 October 2022
from https://phys.org/information/2022-10-collaborative-team-natural-products-sources.html
This doc is topic to copyright. Aside from any honest dealing for the aim of personal research or analysis, no
half could also be reproduced with out the written permission. The content material is supplied for data functions solely.